postgrad - UQ:
undergrad - JCU:
high school - SSHS
Multi-Scale, Spatio-Temporal Analysis of Mammalian Cell Tomograms
PhD Thesis by Andrew Noske
 Introduction:
Introduction:
This page includes an abstract and other information from my PhD thesis, which I undertook at the University of Queensland (UQ) in Brisbane Australia. Due to UQ requirements, I am not allowed to provide a hyperlink to my actual thesis from my site, however I can provide you with a link to an assessment of my thesis (which also includes a breakdown of each chapter), and a copy of the abstract (see below). If you are interested in obtaining a copy of my thesis don't hesitate to e-mail me and I'll provide a hyperlink.
Project Details:
- Thesis Title: ..... Multi-Scale, Spatio-Temporal Analysis of Mammalian Cell Tomograms
- Author: ............ Andrew Noske
- Institute: .......... Institute for Molecular Bioscience (IMB), The University of Queensland (UQ), Brisbane St Lucia Campus
- Primary supervisor: ........ Dr. Brad Marsh
- Associate supervisors: ... Prof. Kevin Burrage and Prof. Mark Ragan
- Date submitted: .............. Feb 2010
Abstract:
The biological, technical and computational aspects of this project collectively focused on using electron tomography (ET) for the high-resolution (10-20 nm) 3D reconstruction of entire insulin-secreting beta cells within islets of Langerhans isolated from mouse pancreata. Islets were cultured overnight to represent either steady-state (non-stimulated) or elevated glucose (stimulated) conditions, prior to fast-freezing, freeze-substitution, plastic embedment and cutting into 250-400 nm thick sections for tomographic imaging using intermediate voltage electron microscopy (EM). 3D images (tomograms) of each section were used to evaluate the performance of the new technical and computational approaches developed, and make biological comparisons of intercellular structure-function. Analysis focused on key compartments/organelles of the insulin-secretory pathway - Golgi apparatus, mitochondria, insulin secretory granules and multi-granular bodies.
To allow the application of ET to entire mammalian cells, several technical limitations were addressed. Since segmenting (delimiting compartments of interest) tomograms manually, represented the major ‘rate-limiting step’ of ET, an interactive approach for 3D segmentation using novel interpolation algorithms (crude smooth, pointwise smooth and spherical interpolation) to iteratively predict the shape of 3D surfaces between user-drawn contours was developed. The performance of these tools in segmenting a range of compartment types was examined, and found to significantly enhance the speed and accuracy of manual segmentation. To better compensate for the physical collapse of plastic sections in the EM, a novel method was developed for estimating section collapse by analyzing approximately spherical organelles. Using this method on mature insulin granules in high-resolution datasets, coupled with measurements from the whole cell reconstructions, section collapse was found to be substantially less (~25%) than the value (40%) previously used to re-scale 3D models. Other new approaches developed to further improve the accuracy and quality of tomograms, included interactive tools for fiducial tracking, and the use of larger gold particles, a ‘reduced second axis’ to account for the missing wedge problem, and deformation grids to account for anisotropic deformation.
As well as affording more efficient and precise mapping of cell ultrastructure in 3D for subsequent quantitative analysis, these developments provided new insights for future automated (hybrid) segmentation pipelines and new computational approaches for improving quality and isotropic accuracy of volumetric image data. The Interpolator and DrawingTools for segmentation, AnalysisTools for estimating section collapse and BeadHelper for tracking fiducial particles, written as plug-ins for the IMOD software package distributed by the University of Colorado, are now being used by the wider ET community with significant positive feedback.
Using the novel approaches developed, four insulin-secreting beta cells - two from the periphery of an islet frozen 1 hr after stimulation with 11 mM glucose, and two from the periphery of another islet under steady-state 5.6 mM glucose conditions - were reconstructed in their entirety in 3D. Quantitative data on the key compartments/organelles provided new information regarding global changes in cellular organization, and enabled robust comparisons of each pair of functionally equivalent cells at unprecedented spatial resolution. Relative differences in the number, dimensions, architecture and distribution of organelles per cubic micron of cellular volume (including mitochondrial branching) reflected differences in the cells’ individual capacity/readiness to respond to secretagogue stimulation. In the two stimulated cells this was reflected by inverse relationships between the number/size of mature granules versus immature granules, the number/size of mitochondria, and the volume of the trans-Golgi network relative to the entire Golgi ribbon. Complementary stereological analysis of whole islets indicated which cells were the most representative under stimulated versus non-stimulated conditions, and revealed a marked natural heterogeneity between cells both within and between individual islets.
Overall, this project led to significant improvements in efficiency and accuracy for segmenting cellular compartments/organelles, and in image quality and accuracy for tomogram computation and reconstruction through use of the newly developed techniques. The improved 3D reconstruction and analysis of pancreatic beta cells in toto in native tissue provided a powerful approach for quantitatively mapping the organelles involved in insulin synthesis/secretion at unprecedented detail, and afforded a level of insight into the complex 3D organization of mammalian cells not previously achieved by any other analytical technique or imaging method.
Keywords:
- electron microscope tomography
- three-dimensional spatial analysis
- interpolation
- automatic segmentation
- beta cell
- mouse pancreas
- insulin secretory pathway
- image processing
- whole cell tomography
About the project:
This project represents four-and-a-half years of work and experiments, not including the first six-months of my Australian Postgraduate Award (APD) which I spent searching for this project! When I moved to UQ I knew I wanted to find a project I felt was meaningful and visionary, and a project I could enjoy and commit to for several years. Although it took six months, the effort paid off when I found Brad Marsh and shortly afterwards I changed my supervisors and (by necessity) shifted from UQ's ITEE department to IMB. What I loved most about this project was the vision to capture a whole cell at high resolution in ET, despite the fact no-one had come even close to capturing that volume of mammalian cell using ET before. This presented a large array of new computational challenges and opportunities to work on scientific visualization tools. It also meant I had to learn cell biology. Due to the Marsh Group belonging to the Cell Biology division of IMB, almost all my presentations were focused almost entirely on "biological insights", with only a passing comment on the new algorithms and technical aspects of my work which formed the foundation of my project - but at the same time I could understand why this was important and why it was also important not to identify myself as having an IT background or being a "computer scientist" in front of cell biologists. It's far better not to label oneself like this and let the people around you eventually realize in their own time that your skill-set and project is multi-disciplinary.
Over the course of my PhD I gave a total of 12 presentations (not including group meetings) including 5 competitively selected talks. Because I loved my project and was able to develop many 3D animations to present my data, delivering talks was probably the favourite part of my PhD - the other being receiving positive feedback on my IMOD plugins. The least favourite part of my work was spending hours upon hours (easily >50% of my whole thesis) doing what I called "monkey work" - i.e. tracking fiducial (tracking black dots) and segmenting data (tracing outlines) and so I'd always like to remind people that the pretty electron tomography models they see represents many hours - in fact often many months of hard work - except in the rare cases where the data is so clean it can be segmented automatically. One disappointment of my project is that my supervisor and I were, at the start of the project, almost sure we'd have time to feature this work on the Australian television science show Catalyst. Alas that dream was all but forgotten due to a couple of crises and political factors I can't mention here. Overall I did very much enjoy my time at IMB, and the many wonderful people in it, and it was only toward the end of my PhD when my scholarship expired that I knew I wanted to try science outside of Australia! My big hope is that the work we did (Brad, myself and many of Brad's other members including Adam Costin, Garry Morgan and Peter Van Der Heide) in developing "whole cell tomography" is continued and within the next few years we see more and more efforts at creating "cellular atlases" of cells in various conditions so that we might better understand not only how the cell works, but better understand diseases such as diabetes!
Send an e-mail to me if you'd like a copy of my thesis or any of my publications (listed here).
Kind regards,
Dr Andrew Noske. (![]() )
)
Selected images from the thesis:
Due to the nature of my thesis, it included many figures - almost 100 in total! To help get you excited I've included a small subset of these images below in (deliberately) low resolution. To see the rest you'll need to e-mail me!
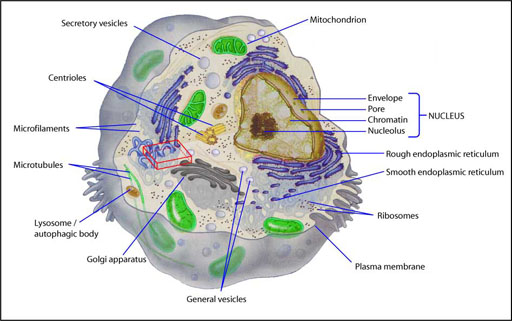
Figure 1.1 Depiction of a mammalian cell showing compartmentalization into functionally and spatially distinct organelles.

Figure 1.4 Insulin structure and molecular synthesis.

Figure 1.14 Improved 3D reconstruction accuracy using a dual-axis tomography approach.

Figure 2.1 Location of the two ‘stimulated’ cells under study within the glucose-stimulated islet of Langerhans

Figure 2.3 Side views of each final whole cell tomogram showing how many sections/tomograms were imaged to reconstruct each cell in toto.

Figure 3.1 Example of the structural complexity of subcellular organization revealed in a high-resolution (~6 nm) tomogram centered on the Golgi region in a mammalian cell

Figure 4.22 Identifying and tracking gold fiducial markers in whole cell sections

Figure 4.21 Result of applying XY deformation grid to the whole cells

Figure 5.6 Methods used to segment and estimate the volume of the main Golgi ribbon

Figure 5.7 3D organization of the Golgi ribbon

Figure 5.10 Quantitative analysis of variations in mitochondrial morphology and length

Figure 5.13 A subset of mitochondria in cell03 reveal novel structure and function

Figure 5.16 Invaginations on the external surface of cell02.

Figure 5.21 General membrane traffic - tubular and vesicular carriers